DNA Extraction of Oly Broodstock
-
SS1_1SS1_2 (not tan)SS1_3SS1_4SS1_5SS1_6SS1_7SS1_8SS1_9SS1_10SS1_11SS1_12SS1_13SS1_14SS1_15SS1_16SS1_17SS1_18SS1_19SS1_20SS3_1SS3_2SS3_5SS3_6
- Found 2 tubes labelled SS1_2. Marked the unextracted one with a tan sticker
- Could not initially find SS3_3 or SS3_4 but found later and reorganized box
- Left in 60degC water bath for 3 hours
- Eluted 50 uL then 100 uL
Primers
- Received the rest of the 2b-RAD primers and resuspended them to 100 uM.
- BC11
- BC12
- HT3
- HT4
- HT5
- HT6
- HT7
- HT8
- Made new 1 uM aliquots of BC2-10 and HT1
- 1 uL stock + 99 uL NFW
- Made new 10 uM aliquots of Lib 1 and Lib 2
- 2.5 uL stock + 22.5 uL NFW
PCR for 2b-RAD MiSeq run
Used 10 samples from 9/28/15 digestion and ligation. Numbers in parentheses are so I know which tube from the ligation each sample is in.
- SS2_6 (5)
- SS2_7 (6)
- SS2_8 (7)
- HC1_5 (8)
- HC1_6 (9)
- HC1_7 (10)
- HC1_8 (11)
- NF1_4 (14)
- NF1_5 (15)
- NF1_6 (16)
Master Mix
|
1x
|
10.5x
|
|
|
NFW
|
23 uL
|
241.5
|
|
10 mM (each) dNTPS
|
2
|
21
|
|
10 uM ILL-Lib1
|
2
|
21
|
|
10 uM ILL-Lib2
|
2
|
21
|
|
5X Q5 buffer
|
20
|
210
|
|
Q5 Taq polymerase
|
1
|
10.5
|
Added 50 uL of MM to 40 uL of ligation product. Added 5 uL of 1uM BC and HT to each sample, using BC1-10 and HT1.
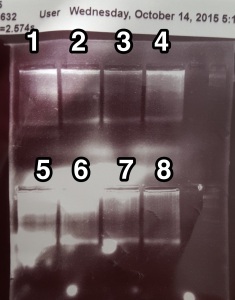
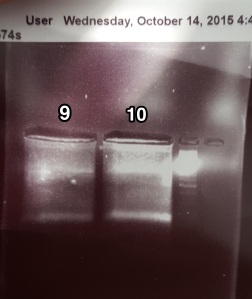
Run out gel of 2b-RAD PCR product
This time, set up a medium gel for 8 of the samples and a small gel for the other 2. Made a 2% TAE gel, and after running it at 110V for 1 hour let the gels soak in water with 6 uL of EtBr to increase staining. PCR used 19 cycles.
I think I should make sure to mix in the EtBr when doing the post staining as that is likely why there are bright white sections on the gels. It was still much easier to see the bands though. Cut out the bands and left them in 1.5 mL tubes in 4degC for gel extraction on Friday.


Pingback: Friday 10/16/15 | Getting My Genes Wet